Increasingly, researchers are turning to big data sets and large-scale computational models to support policymakers during epidemic emergencies. Image courtesy of gettyimages/ktsimage.
Latest News
November 9, 2018
When infectious diseases such as the Ebola and Zika viruses threaten to spread, epidemiologists look for ways to forecast the trajectory of an outbreak as quickly as possible to assist the public health policy response. Increasingly, researchers are turning to big data sets and large-scale computational models to support policymakers during epidemic emergencies. Mathematical models can generate forecasts of the progression of epidemics based on a variety of variables and potential actions taken to counter their spread.
One example is the Research and Policy for Infectious Disease Dynamics (RAPIDD) program coordinated by the Fogarty International Center at the National Institutes of Health (NIH). In 2015, RAPIDD ran an Ebola modeling exercise in which eight teams from U.S., U.K. and Canadian universities and several U.S. government agencies were tasked with predicting when Ebola would peak in Liberia and how it would progress between September and December 2015. Each team was given four different Ebola scenarios to model, each with different levels of containment. The teams were free to choose the parameters they would add to their model. The results were presented and compared at a two-day forum.
“Ebola was the first case of developing such a model anywhere in Africa,” says Cecile Viboud, a senior scientist at the Fogarty Center. The models contained detail about the population’s household makeups, how people traveled, data on environmental factors, as well as fine-grained detail about the disease transmission process. “We had them predict a few weeks ahead the trajectory and watched how well their models worked,” she says. “The idea is to try to learn for the next time with better models and to have better coordination between teams when the next crisis arises.”
All these models are run on high-performance computer clusters. A simulation is just one representation of the results you could get, Viboud says, so doing hundreds of simulations with slightly tweaked parameters takes time, even with a big cluster. All of the modeling teams involved have access to high-performance clusters to run their models, and software has been developed to handle the management and analysis of epidemic simulations stemming from large-scale computational models. For instance, a platform called epiDMS was developed to address the need to generate, search, visualize and analyze large volumes of epidemic simulation data and observations during the progression of an epidemic.
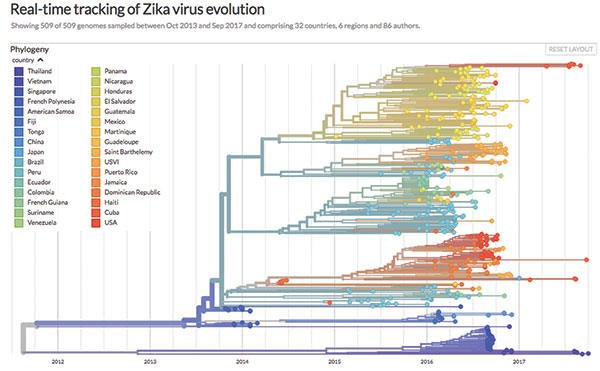
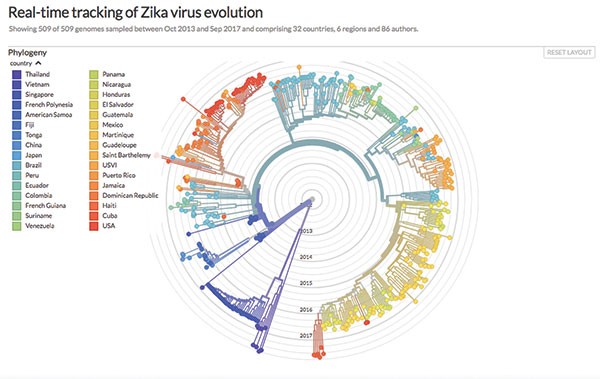
Viboud says that one research group RAPIDD has worked with is led by Alessandro Vespignani, a professor at Northeastern University, whose team also developed models and interactive maps to make projections about the spread and impact of the Zika virus in 2017. According to a description on the Northeastern website, his team of 14 researchers “uses large-scale computational epidemic models that integrate socio-demographic and travel data of target populations along with simulations of infection transmission among millions of individuals to reconstruct disease spread in the past and project it into the future.”
A sister effort to RAPIDD at NIH is called The Models of Infectious Disease Agent Study (MIDAS), a collaboration of research and informatics groups to develop computational models of the interactions between infectious agents and their hosts, disease spread, prediction systems and response strategies. MIDAS says it has produced a number of software packages to help local, state and federal public health officials prepare for and respond to infectious disease emergencies.
Studying Air Travel
Another approach to studying the spread of infectious diseases is to look at air travel. Funded by the National Science Foundation, the VIPRA Project is a multi-university effort to analyze new strategies for reducing the risk of spread of viral infections through air travel. (VIPRA stands for Viral Infection Propagation Through Air Travel.)
When the Ebola outbreak in Africa happened, some people called for travel bans from affected countries, notes Ashok Srinivasan, a professor in the Department of Computer Science at the University of West Florida and a lead investigator of the VIPRA Project. Travel bans, however, have a big impact, not just on the spread of the infection, but on local economies and people’s lives, he adds. “Can we take some actions that are not as drastic as banning air travel but at the same time would reduce the likelihood of infection spreading? That is our goal — to identify procedural or policy steps that could be taken to decrease the likelihood of infection spreading without disrupting air travel.”
The researchers are integrating models of how people move and interact during air travel with a computational infrastructure that is designed for simulation-based policy analysis on some of the most powerful supercomputers in the world, including Blue Waters at the National Center for Supercomputing Applications at the University of Illinois. (Blue Waters uses hundreds of thousands of computational cores to achieve peak performance of more than 13 quadrillion calculations per second.)
Srinivasan explains how the VIPRA Project takes advantage of Blue Waters. “We parameterize the sources of uncertainty and perform simulations to cover the range of uncertainties,” he says. For instance, they look at how long it takes to board a plane and stow luggage. Once those variations are taken into account, the number of scenarios becomes very large and a sequential computation cannot solve it, he says. “This approach leads to high computational cost, which is handled through massive parallelization on Blue Waters.”
Srinivasan says VIPRA’s results show potential for substantial reduction in Ebola spread by changing current boarding and disembarkation procedures. Among their findings:
- Random boarding leads to lower risk of infection spread;
- Boarding has a higher impact than deplaning; and
- Smaller planes are better than larger ones in terms of lower risk of infection spread.
The VIPRA team has worked to fine-tune its simulations to get better run times and has been able to take them from 20 minutes to less than 1 minute, Srinivasan says. “Blue Waters provides us access to a large number of processors so that, if a new infection comes, we will be up and running really quickly.”
Genomic Epidemiology
Genomic epidemiology is a field in which whole genome sequencing is used to investigate key infectious disease-causing microbes with a goal of understanding how they spread and cause illness. A Canadian open source project, the Integrated Rapid Infectious Disease Analysis (IRIDA) platform was developed to support real-time infectious disease outbreak investigations using whole genome sequencing data. IRIDA is designed for use in public health, food safety and clinical microbiology labs.
One of the principal investigators, William Hsiao, a senior scientist at the British Columbia Centre for Disease Control’s Public Health Laboratory, says the IRIDA platform was set up to allow DNA sequence data from disease-causing bacteria to be shared with other researchers. “By achieving more timely data sharing and allowing more people to do analysis, we are streamlining the process of infectious disease outbreak investigations,” he says.
IRIDA is taking advantage of Cedar, the new advanced research computing system at Simon Fraser University in Burnaby, B.C., which is the most powerful academic supercomputer in Canada.
Analyzing and processing the DNA sequencing is both memory- and CPU-intensive, Hsiao explains. “Having a large cluster means we can be scalable and can process more data in parallel. Previously available clusters were designed more for CPU-intensive tasks commonly used in physics and math, but bioinformatics work requires more memory,” he says. “The new Cedar cluster has nodes that are designed to facilitate high-memory analyses that are quite common in bioinformatics.”
The IRIDA project leaders are working toward making the software platform more portable with a specific goal of making it cloud-compatible. “That way, people could easily deploy the platform in the commercial cloud without having to have a dedicated cluster or dedicated personnel to install the software,” Hsiao says. “We want the IRIDA platform to be compatible with different high-performance computing environments, and we are working toward that.”
For mathematical models to have a greater impact on infectious disease response, it is important to have modelers embedded in public health settings, says the Fogarty Center’s Viboud. She adds that, although modelers are interested in research questions and in creating the best model possible, there is an issue of timeliness. “If you want to make a prediction about public health, you have to be quick and applied, so you have to find the right interface between modelers and public health experts,” she adds. “That is a dialog that is still happening.”
More Dell EMC Coverage
For More Info
Subscribe to our FREE magazine, FREE email newsletters or both!
Latest News